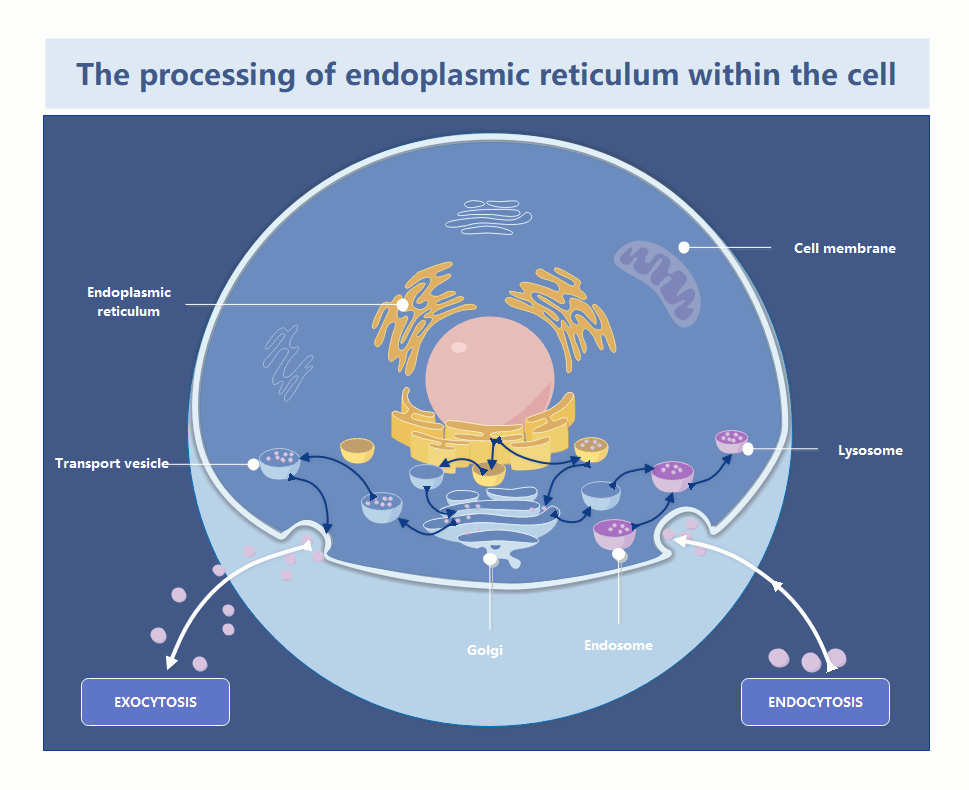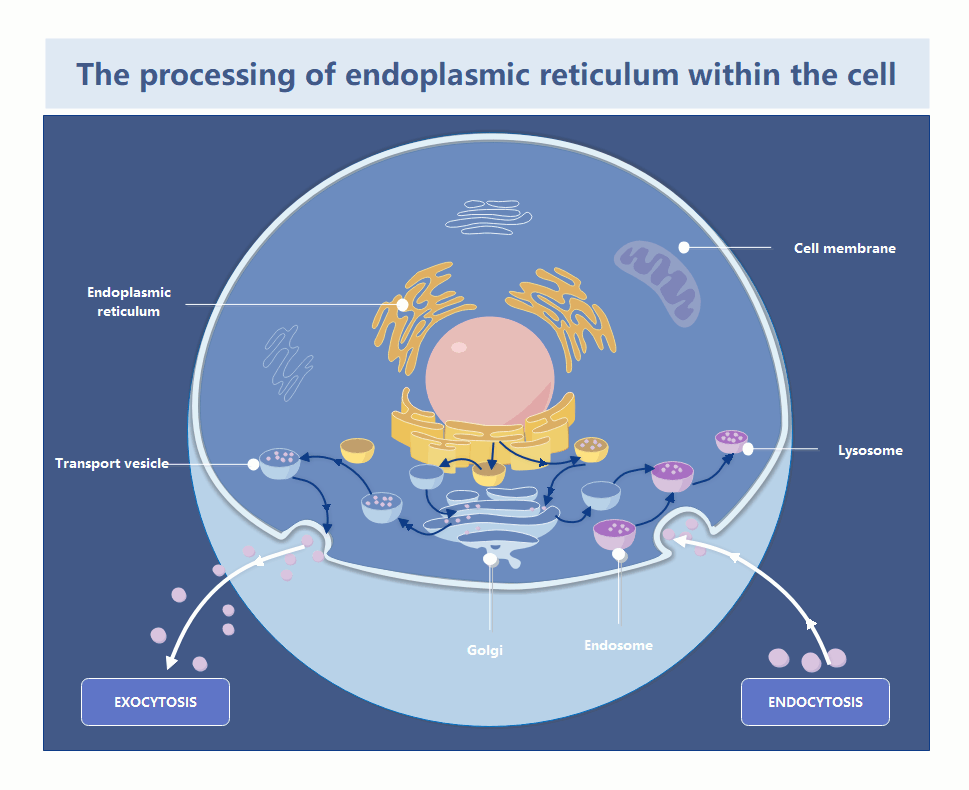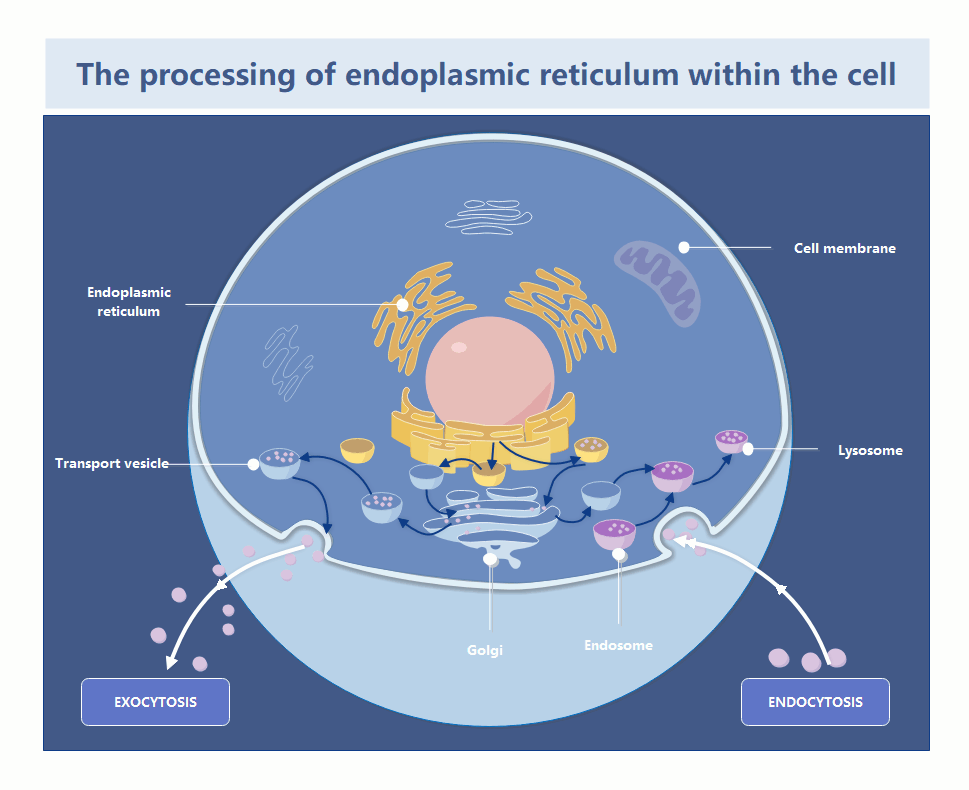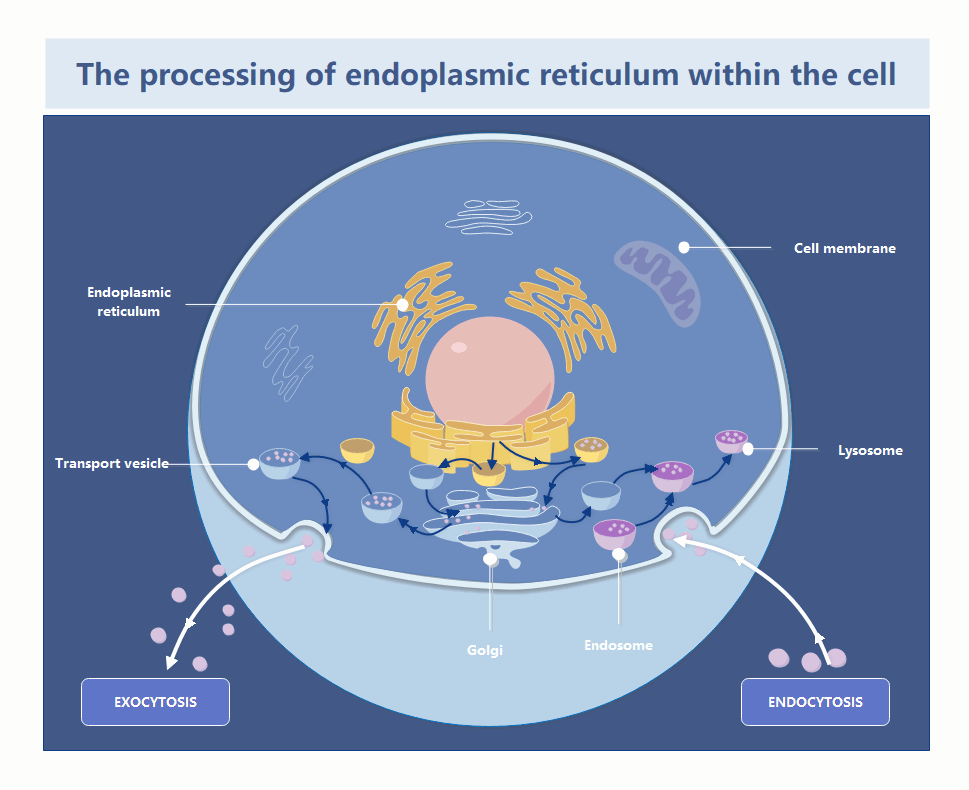
How Cells Remove Damaged Endoplasmic Reticulum
Biologists just mapped the cellular equivalent of a garbage collection routine for the endoplasmic reticulum (ER). While the press release smells like academic curiosity, the actual mechanism—ER-phagy—is essentially a hardware-level system purge that prevents cellular “kernel panics” and systemic failure.
The Tech TL;DR:
- Mechanism: Discovery of specific receptor-mediated autophagy (ER-phagy) that targets damaged ER membranes for degradation.
- Systemic Impact: Failure in this “cleanup” process correlates directly with neurodegenerative “bit rot” and metabolic crashes.
- Enterprise Parallel: This is the biological version of automated container orchestration and resource reclamation to prevent memory leaks.
In the world of systems architecture, we talk about memory leaks and zombie processes. In the cytosol, the endoplasmic reticulum is the primary factory for protein folding and lipid synthesis. When the ER becomes “damaged”—whether through oxidative stress or misfolded protein accumulation—it doesn’t just sit there; it becomes a toxic bottleneck. If the cell cannot prune these defective segments, the entire organelle enters a state of chronic stress, triggering the Unfolded Protein Response (UPR), which, if left unchecked, initiates a programmed shutdown (apoptosis).
The recent findings, detailed in the foundational research published via Nature and summarized by Phys.org, clarify the precise signaling pathway used to flag these damaged zones. This isn’t a random vacuuming process; it is a targeted, receptor-driven operation. By identifying the specific proteins that act as “tags” for the autophagosome, researchers have essentially reverse-engineered the cell’s internal decommissioning API.
For those of us managing high-availability clusters, the analogy is clear: the cell is running a continuous integration/continuous deployment (CI/CD) cycle where the “damaged” ER is the failing build that must be rolled back or purged to maintain system stability. When this process fails, we notice the biological equivalent of a cascading failure across the network—manifesting as Parkinson’s or Alzheimer’s.
The “Garbage Collection” Logic: ER-phagy vs. Systemic Decay
To understand the gravity of this, we have to gaze at the “blast radius” of ER dysfunction. When the ER fails to clear damaged sections, the cell experiences a massive increase in latency for protein synthesis. This isn’t just a performance hit; it’s a critical failure. In a computing environment, this would be like a memory leak in a C++ application that eventually consumes all available RAM, forcing the OOM (Out of Memory) killer to terminate the process.

“The precision of ER-phagy is the only thing preventing the cytosol from becoming a graveyard of misfolded proteins. If the receptor-tagging mechanism lags, the cell doesn’t just unhurried down—it crashes.” — Dr. Aris Thorne, Lead Computational Biologist at the Synthetic Genomics Initiative.
From a technical standpoint, the “tagging” mechanism functions like a metadata header in a packet. The cell identifies a “corrupt” segment of the ER, attaches a specific protein receptor (the metadata), and the autophagosome (the cleanup script) recognizes that header and encapsulates the segment for enzymatic digestion. This is high-precision resource management. If you’re running a legacy infrastructure and seeing similar patterns of systemic decay, it’s time to bring in enterprise IT infrastructure auditors to map your dependencies before the “biological” rot sets in.
Simulating Cellular Cleanup: The Logic Gate
While we can’t write Python to fix a mitochondria, we can model the logic of receptor-mediated cleanup. If we were to represent the ER-phagy trigger as a conditional check in a resource management script, it would look something like this:
# Pseudo-code for ER-phagy Resource Reclamation def cellular_cleanup_cycle(er_segment): # Check for protein misfolding or membrane damage (The 'Triage' phase) if er_segment.status == "DAMAGED" or er_segment.protein_load > THRESHOLD: # Attach receptor tag (The 'Flagging' phase) er_segment.apply_tag("FAM20C_RECEPTOR") # Trigger Autophagosome recruitment try: autophagosome.encapsulate(er_segment) lysosome.digest(autophagosome) print("Resource reclaimed: ER segment purged.") except SystemicFailure: trigger_apoptosis() # System shutdown to prevent contagion else: continue_protein_synthesis() The Biological Stack vs. Synthetic Alternatives
The discovery of these specific ER-phagy pathways opens the door to “bio-patching.” We are moving toward an era where we don’t just treat the symptom (the disease) but optimize the underlying “hardware” (the cell). Below is a comparison of how this natural process stacks up against current synthetic interventions.
| Feature | Natural ER-phagy | Little Molecule Inhibitors | CRISPR-based Gene Editing |
|---|---|---|---|
| Precision | High (Receptor-specific) | Low (Systemic effect) | Very High (Sequence-specific) |
| Latency | Real-time / Dynamic | Delayed (Pharmacokinetic) | Permanent (One-time deploy) |
| Risk | Low (Endogenous) | Moderate (Off-target toxicity) | High (Off-target mutations) |
| Scalability | Cell-wide | Dose-dependent | Cell-line specific |
This “biological stack” is currently being analyzed by firms specializing in specialized biotech software development to create digital twins of cellular pathways. By simulating these receptor-mediated cleanses, researchers can predict which drugs will “upregulate” the cleanup process, essentially overclocking the cell’s ability to remove waste.
The “Bio-Security” Bottleneck
As we decode these pathways, we encounter a new kind of vulnerability. If a pathogen or a synthetic agent can “spoof” the ER-phagy tags, it could trick the cell into digesting its own healthy organelles—a biological Denial of Service (DoS) attack. This is why the intersection of AI and cybersecurity is no longer just about protecting servers, but about protecting the biological data that defines us. As enterprise adoption of AI-driven drug discovery scales, the demand for certified cybersecurity consultants who understand the sensitivity of genomic data is skyrocketing.
The research is backed by significant public funding and academic grants, typically managed through institutions like the NIH or European Research Council, ensuring that the foundational “source code” of these discoveries remains in the public domain rather than locked behind a proprietary corporate firewall. This open-source approach to biology allows for faster iteration and peer-review, mirroring the way the Linux kernel evolved into the world’s most stable OS.
We are witnessing the transition from “descriptive” biology to “algorithmic” biology. We aren’t just observing that cells clean themselves; we are documenting the API calls they use to do it. The trajectory is clear: once we master the “garbage collection” of the cell, we can potentially halt the degradation of the human brain. But until then, we’ll keep optimizing our Kubernetes clusters and hoping our own biological hardware doesn’t hit a critical memory leak.
Disclaimer: The technical analyses and security protocols detailed in this article are for informational purposes only. Always consult with certified IT and cybersecurity professionals before altering enterprise networks or handling sensitive data.
