Whole genome sequencing (WGS) is rapidly becoming a feasible and valuable tool in routine cancer diagnostics, offering more comprehensive insights than traditional DNA panel testing, according to a new analysis of nearly 900 patients. The study, conducted between January 2021 and November 2022, demonstrates WGS’s ability to not only identify the tissue of origin for cancers of unknown primary (CUP) but also to reveal actionable biomarkers for treatment in a significant majority of cases.
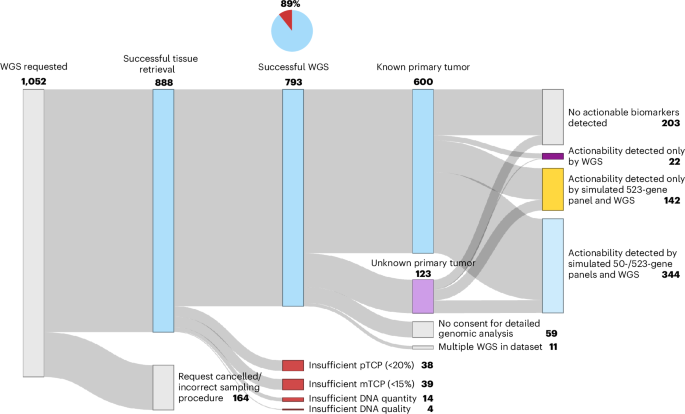
Researchers successfully generated diagnostic sequencing reports for 89% of the 888 tissue samples deemed suitable for WGS. While success rates were lower for cytology samples (57%) compared to surgical specimens or biopsies (91%), the median turnaround time from sample reception to reporting was a swift six working days. This speed is crucial for informing timely treatment decisions.
The analysis, which included patients with various cancer types – notably non-small cell lung cancer (NSCLC) at 23% and CUP at 16% – revealed that 73% of the 723 patients with available data harbored at least one potentially actionable biomarker. Importantly, WGS detected actionable biomarkers in a greater proportion of patients than simulated testing using commonly used 50-gene and 523-gene panels, identifying additional actionable biomarkers in 8% of cases where comprehensive panel testing would have been sufficient.
For patients with CUP, WGS successfully predicted the tissue of origin with a high degree of confidence (confidence score ≥0.80) in 49% of cases, and in an additional 14% when combined with existing clinical and pathological findings. This led to a definitive tumor-type diagnosis for 63% of CUP patients, a significant improvement over standard diagnostic methods. The CUPPA algorithm, used in the analysis, is proving instrumental in resolving these challenging cases.
The study also highlighted the potential of WGS to identify potentially germline variants (PGVs) – genetic changes inherited from parents – that could have implications for patients and their families. PGVs were identified in 6.5% of patients, with about half of these not previously detected through routine diagnostics. Patients were then referred for genetic counseling and further germline analysis.
Overall clinical utility was observed in 41% of all patients who underwent WGS, with clinically relevant results including the identification of biomarkers indicating reimbursed treatment, a solved or clarified diagnosis, or a clinically relevant PGV. Patients with at least one actionable biomarker who received biomarker-informed therapy after WGS demonstrated improved overall survival compared to those with actionable alterations who did not receive such treatment (hazard ratio = 0.78). This benefit was most pronounced in patients without prior systemic therapy.
Researchers found that WGS provided clinically valuable insights for 35% of patients with known primary tumor diagnoses and 67% of those with CUP. The study underscores the growing role of comprehensive genomic sequencing in personalizing cancer treatment and improving patient outcomes, particularly in cases where standard diagnostic approaches fall short.
Leave a Reply